A08 Evolution of transcription factor mutations in myeloid leukemias
Recently, both inherited and acquired mutations of the transcription factor GATA2 were discovered in a variety of myeloid malignancies including AML, MDS and CML. To further elucidate the role of such mutations in leukemia, we will assess their transforming potential in primary murine and human hematopoietic stem cells and in the AML cell line HL60.
Moreover, we will study the impact of GATA2 mutations on DNA-binding, protein-interactions and target-gene regulation. Finally, we will analyze, if the genetic background potentially provides a fertile ground for the acquisition of specific mutations.
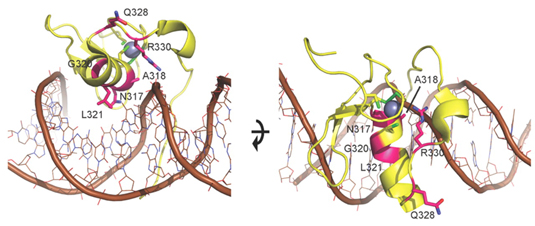
Figure Legend: Model of N-terminal zinc finger of GATA2. Model of ZF1 (yellow ribbon with brown DNA and gray zinc ion), based on the crystal structure of the DNA complex of the highly homologous zinc finger of GATA338 (PDB accession 3DFV). Mutated residues (magenta) are annotated and displayed with side chains. The mutations cluster at the DNA binding side of ZF1, suggesting they perturb DNA binding. Based on the homology model, N317, A318, L321, and R330 are directly implicated in DNA binding, so mutations in these residues probably alter the affinity to DNA or prevent DNA binding. G320 is important for proper attachment of an adjacent β-hairpin loop that provides additional DNA binding contacts. Q328 is not directly involved in DNA binding. However, Q328P could either affect the backbone fold of ZF1 and indirectly alter DNA binding by perturbing the adjacent R330, or alternatively is involved in interaction with other domains such as ZF2. All in all, the location of the mutations suggests that they influence DNA binding of ZF1. From Philipp A. Greif et al. Blood 2012:120:395-403

Dr. Philipp A. GreifMedizinische Klinik und Poliklinik III, Ludwig-Maximilians-Universität München +49 (0)89 4400-43982 |

Prof. Dr. Wolfgang HiddemannMedizinische Klinik und Poliklinik III, Ludwig-Maximilians-Universität München +49 (0)89 4400-72551 |

