A06 Clinical and functional relevance of subclonal driver mutations in acute myeloid leukemia
Many patients with acute myeloid leukemia (AML) carry driver mutations that are not present in the entire tumor cell population, but only in a subclone. In this project, we plan to use highly sensitive methods to search for relapse-associated subclones in pretreatment patient specimens. We will use single-cell RNA sequencing to study phenotypic differences between genetically defined subclones. Finally, we will use a murine xenograft model to understand the functional consequences induced by the gain of a subclonal driver mutation in genetically defined patient-derived AML cells.
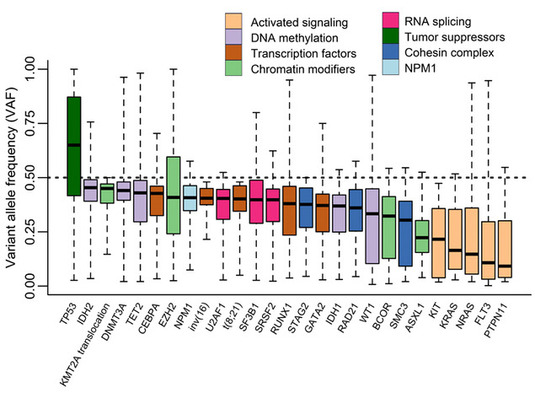
Figure legend: Variant allele frequencies (VAFs) of common driver gene mutations in >600 patients with acute myeloid leukemia (AML). Variants with VAFs near 50% (left side) are present in almost all leukemic cells and probably originate from the founding clone. Variants with lower allele frequencies (right side) may be present in a subpopulation (subclone) of the leukemia cells only. Our aim is to use single-cell techniques to uncover the clonal architecture of AML in individual patients, and to understand the functional consequences of such subclonal mutations.

Dr. Klaus MetzelerMedizinische Klinik und Poliklinik III, Ludwig-Maximilians-Universität München +49 (0)89 4400-72551 |

